Discovering Conserved Spatial Ecotypes Across Multiple Spatial Transcriptomics Samples
Source:vignettes/Integration.Rmd
Integration.RmdOverview
In this tutorial, we will illustrate how to identify conserved spatial ecotypes (SEs) across multiple single-cell spatial transcriptomics (ST) datasets using the SpatialEcoTyper framework.
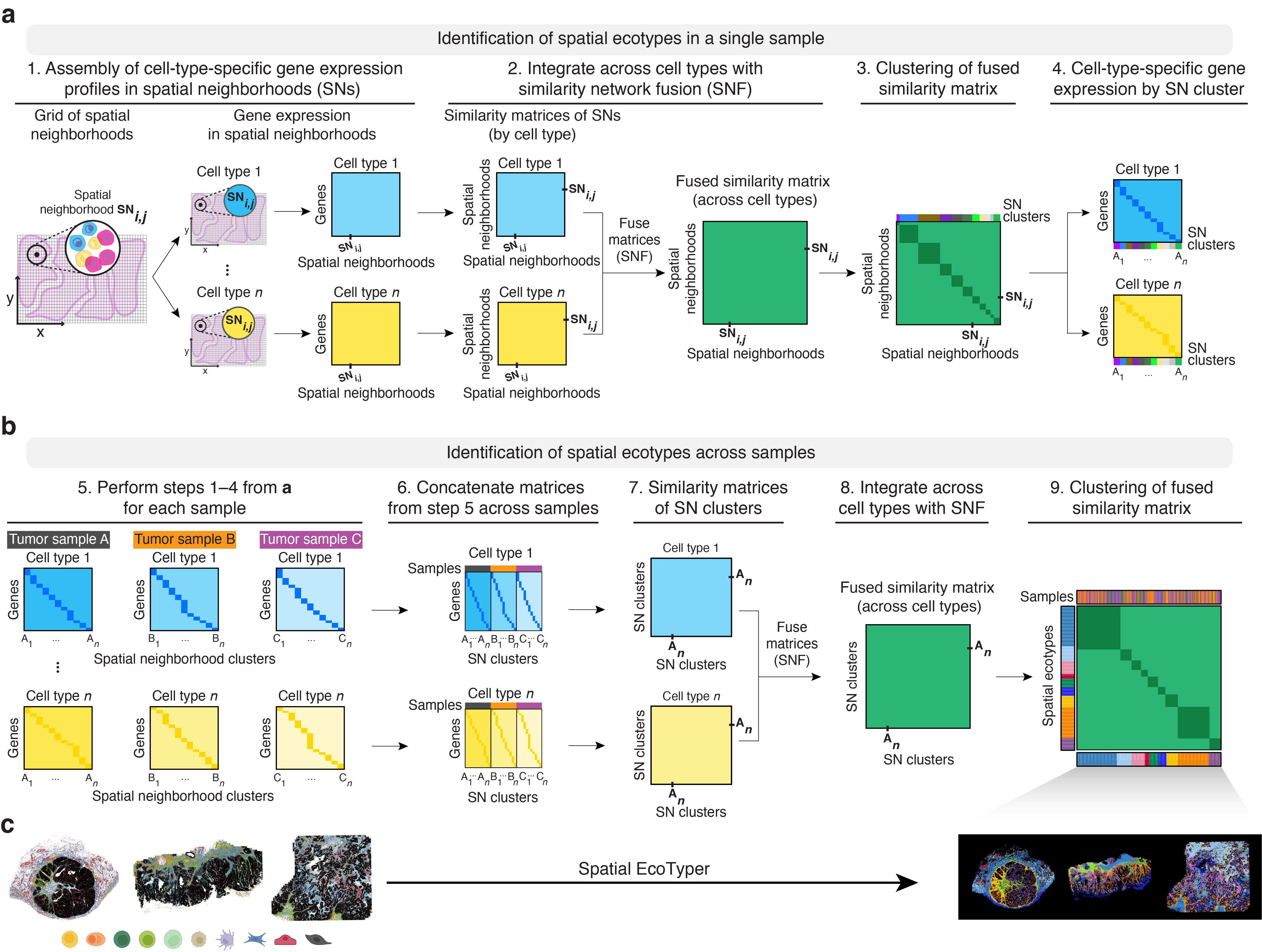
The analysis is performed in two stages:
- Determination of sample-level spatial clusters (Steps 1-4): Spatial neighborhood clusters from each single-cell ST sample can be identified using the SpatialEcoTyper function. Check Tutorial 1 for details.
- Identification of conserved spatial ecotypes (Steps 5-9): Spatial neighborhood clusters discovered from individual single-cell ST samples are represented by gene expression profiles (GEPs) of their associated cell states, and clusters with similar GEPs are aggregated across samples to identify conserved SEs.
Load required packages for this vignette
suppressPackageStartupMessages(library(dplyr))
suppressPackageStartupMessages(library(ggplot2))
suppressPackageStartupMessages(library(parallel))
suppressPackageStartupMessages(library(Seurat))
suppressPackageStartupMessages(library(data.table))
suppressPackageStartupMessages(library(NMF))
suppressPackageStartupMessages(library(ComplexHeatmap))
library(SpatialEcoTyper)Loading data
For the demonstration of the integrative analysis of multiple single-cell ST samples, we have selected a subset of regions from a melanoma sample and a colon cancer sample profiled by Vizgen MERSCOPE.
The melanoma sample includes spatial expression data for 500 genes across 27,907 cells, while the colon cancer sample contains data for 38,080 cells. In both samples, cells were categorized into ten distinct cell types: B cells, CD4 T cells, CD8 T cells, NK cells, plasma cells, macrophages, dendritic cells (DC), fibroblasts, endothelial cells, and cancer cells. Cancer cells are excluded from this demonstration to reduce processing time.
All cells are grouped into four spatial regions: tumor, inner margin, outer margin, and stroma. The tumor (tumor and inner margin) and stroma regions (stroma. and outer margin) are defined based on the density of cancer cells, as described in the CytoSPACE paper. The inner and outer margins are defined as regions extending 250 μm inside and outside the tumor boundaries, respectively.
First, download the demo data and load them into R. You can use
read.table for small files or
data.table::fread for larger files.
# Load single-cell gene expression data. Rows represent gene names and columns represent cell IDs
scdata1 <- fread("https://spatialecotyper.stanford.edu/inc/inc.public.vignettes.php?file=Melanoma1_subset_counts.tsv.gz",
sep = "\t", header = TRUE, data.table = FALSE)
rownames(scdata1) <- scdata1[, 1] # Setting the first column as row names
scdata1 <- as.matrix(scdata1[, -1]) # Dropping first column
scdata2 <- fread("https://spatialecotyper.stanford.edu/inc/inc.public.vignettes.php?file=CRC2_subset_counts.tsv.gz",
sep = "\t", header = TRUE, data.table = FALSE)
rownames(scdata2) <- scdata2[, 1] # Setting the first column as row names
scdata2 <- as.matrix(scdata2[, -1]) # Dropping first column
head(scdata1[,1:5])## HumanMelanomaPatient1__cell_3655 HumanMelanomaPatient1__cell_3657
## PDK4 0 1
## TNFRSF17 0 0
## ICAM3 0 0
## FAP 1 0
## GZMB 0 0
## TSC2 0 0
## HumanMelanomaPatient1__cell_3658 HumanMelanomaPatient1__cell_3660
## PDK4 1 0
## TNFRSF17 0 0
## ICAM3 0 0
## FAP 0 0
## GZMB 0 0
## TSC2 0 0
## HumanMelanomaPatient1__cell_3661
## PDK4 0
## TNFRSF17 0
## ICAM3 0
## FAP 0
## GZMB 0
## TSC2 0
# head(scdata2[,1:5])
## Normalize the data as needed
normdata1 <- NormalizeData(scdata1)
normdata2 <- NormalizeData(scdata2)Start with SpatialEcoTyper results from individual single-cell ST samples
The IntegrateSpatialEcoTyper
function provides a convenient way to integrate Spatial EcoTyper results
across samples or cancer types to identify conserved spatial ecotypes.
This function assumes that SpatialEcoTyper has been
run on each sample using consistent settings. Before proceeding, please
review the full manual of the IntegrateSpatialEcoTyper
function here.
url <- "https://spatialecotyper.stanford.edu/inc/inc.public.vignettes.php?file=CRC2_subset_SpatialEcoTyper_results.rds"
download.file(url, destfile = "CRC2_subset_SpatialEcoTyper_results.rds", mode = "wb")
url <- "https://spatialecotyper.stanford.edu/inc/inc.public.vignettes.php?file=Melanoma1_subset_SpatialEcoTyper_results.rds"
download.file(url, destfile = "Melanoma1_subset_SpatialEcoTyper_results.rds", mode = "wb")Note: If you are using Seurat v4 (rather than Seurat v5), please follow Tutorial 1 to generate Spatial EcoTyper results for the two demo samples before testing the scripts below.
data_list <- list(SKCM = normdata1, CRC = normdata2)
SpatialEcoTyper_list <- list(SKCM = readRDS("Melanoma1_subset_SpatialEcoTyper_results.rds"),
CRC = readRDS("CRC2_subset_SpatialEcoTyper_results.rds"))
IntegrateSpatialEcoTyper(SpatialEcoTyper_list, data_list,
outdir = "SpatialEcoTyper_results/",
normalization.method = "None",
nfeatures = 300, # Number of top variable genes to use
nmf_ranks = 4:15, # Number of clusters to test
nrun.per.rank = 10, # recommend 30 or higher for robust results
ncores = 16)Choosing Louvain clustering resolution: Louvain clustering was applied to group spatial neighborhoods into clusters within each sample, which improves computational efficiency for integrative analysis of samples. We recommend using as high a clustering resolution as feasible (30 or higher) to better preserve fine-grained biological structure and ensure robustness to parameter variation.
Choosing the optimal number of SEs: NMF clustering
is used to identify conserved spatial ecotypes across samples. To
determine the optimal number of SEs, users can specify the
nmf_ranks and nrun.per.rank arguments to
evaluate multiple candidate ranks, each with a defined number of runs
(we recommend ≥30 runs per rank to ensure robustness, as NMF uses random
initialization). For each rank, the cophenetic coefficient is computed
to assess clustering stability. This metric ranges from 0 to 1, with
higher values indicating greater stability. Typically, the cophenetic
coefficient exhibits a multi-modal pattern across ranks, and we selected
the optimal number of SEs as the highest rank at which the cophenetic
coefficient exceeds 0.95 (controlled by the min.coph
argument) and is followed by the largest subsequent decrease, indicating
a transition to less stable solutions.
Balancing predefined regions: If predefined tissue
regions are available (e.g., pathologist annotations of tumor and
adjacent stromal regions), you can use the Region and
downsample.by.region argument to balance spatial
neighborhoods across regions. When enabled, the method downsamples
neighborhoods within each region to ensure comparable representation and
to weight regions equally during integration.
Start with raw single-cell ST data from multiple samples
The MultiSpatialEcoTyper function provides an end-to-end pipeline for the integrative analysis of single-cell spatial transcriptomics data across multiple samples. While this streamlined workflow is convenient, it can be computationally intensive, as single-sample Spatial EcoTyper analysis is the most memory- and time-demanding step. Therefore, we recommend first analyzing each sample individually using the SpatialEcoTyper function, as demonstrated in Tutorial 1. The resulting outputs can then be integrated across samples using the IntegrateSpatialEcoTyper function, as described above.
First, load single-cell gene expression data and meta data.
# Load single-cell gene expression data. Rows represent gene names and columns represent cell IDs
scdata1 <- fread("https://spatialecotyper.stanford.edu/inc/inc.public.vignettes.php?file=Melanoma1_subset_counts.tsv.gz",
sep = "\t", header = TRUE, data.table = FALSE)
rownames(scdata1) <- scdata1[, 1] # Setting the first column as row names
scdata1 <- as.matrix(scdata1[, -1]) # Dropping first column
scdata2 <- fread("https://spatialecotyper.stanford.edu/inc/inc.public.vignettes.php?file=CRC2_subset_counts.tsv.gz",
sep = "\t", header = TRUE, data.table = FALSE)
rownames(scdata2) <- scdata2[, 1] # Setting the first column as row names
scdata2 <- as.matrix(scdata2[, -1]) # Dropping first column
head(scdata1[,1:5])## HumanMelanomaPatient1__cell_3655 HumanMelanomaPatient1__cell_3657
## PDK4 0 1
## TNFRSF17 0 0
## ICAM3 0 0
## FAP 1 0
## GZMB 0 0
## TSC2 0 0
## HumanMelanomaPatient1__cell_3658 HumanMelanomaPatient1__cell_3660
## PDK4 1 0
## TNFRSF17 0 0
## ICAM3 0 0
## FAP 0 0
## GZMB 0 0
## TSC2 0 0
## HumanMelanomaPatient1__cell_3661
## PDK4 0
## TNFRSF17 0
## ICAM3 0
## FAP 0
## GZMB 0
## TSC2 0
# head(scdata2[,1:5])
# Load single-cell metadata
# Row names should be cell ids. Required columns: X, Y, CellType;
# Recommend a 'Region' column if region annotations are available
scmeta1 <- read.table("https://spatialecotyper.stanford.edu/inc/inc.public.vignettes.php?file=Melanoma1_subset_scmeta.tsv",
sep = "\t", header = TRUE, row.names = 1)
scmeta2 <- read.table("https://spatialecotyper.stanford.edu/inc/inc.public.vignettes.php?file=CRC2_subset_scmeta.tsv",
sep = "\t", header = TRUE, row.names = 1)
head(scmeta1[, c("X", "Y", "CellType", "Region")])## X Y CellType Region
## HumanMelanomaPatient1__cell_3655 1894.706 -6367.766 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3657 1942.480 -6369.602 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3658 1963.007 -6374.026 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3660 1981.600 -6372.266 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3661 1742.939 -6374.851 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3663 1921.683 -6383.309 Fibroblast Stroma## X Y CellType Region
## HumanColonCancerPatient2__cell_1085 5057.912 -1721.871 Macrophage Inner margin
## HumanColonCancerPatient2__cell_1205 5035.339 -1783.352 Macrophage Inner margin
## HumanColonCancerPatient2__cell_1312 5012.093 -1908.390 Macrophage Inner margin
## HumanColonCancerPatient2__cell_1625 5073.743 -1861.740 Macrophage Inner margin
## HumanColonCancerPatient2__cell_1629 5030.235 -1867.441 Macrophage Inner margin
## HumanColonCancerPatient2__cell_1639 5020.448 -1882.751 Macrophage Inner margin
# Make sure row names of the meta data match column names of the gene expression matrix
scdata1 <- scdata1[, match(rownames(scmeta1), colnames(scdata1))]
scdata2 <- scdata2[, match(rownames(scmeta2), colnames(scdata2))]
## Normalize the data as needed
normdata1 <- NormalizeData(scdata1)
normdata2 <- NormalizeData(scdata2)Then, you can submit the job and wait.
data_list <- list(SKCM = normdata1, CRC = normdata2)
metadata_list <- list(SKCM = scmeta1, CRC = scmeta2)
MultiSpatialEcoTyper(data_list, metadata_list,
normalization.method = "None", # Normalization method
nmf_ranks = 10, # Number of clusters
nrun.per.rank = 30, # Recommend 30 or higher for robust results
ncores = 2)Optimizing memory usage
The similarity network fusion (Step 2 of SpatialEcoTyper) is the most computationally demanding step, and its runtime increases with the number of spatial neighborhoods (SNs) per sample.
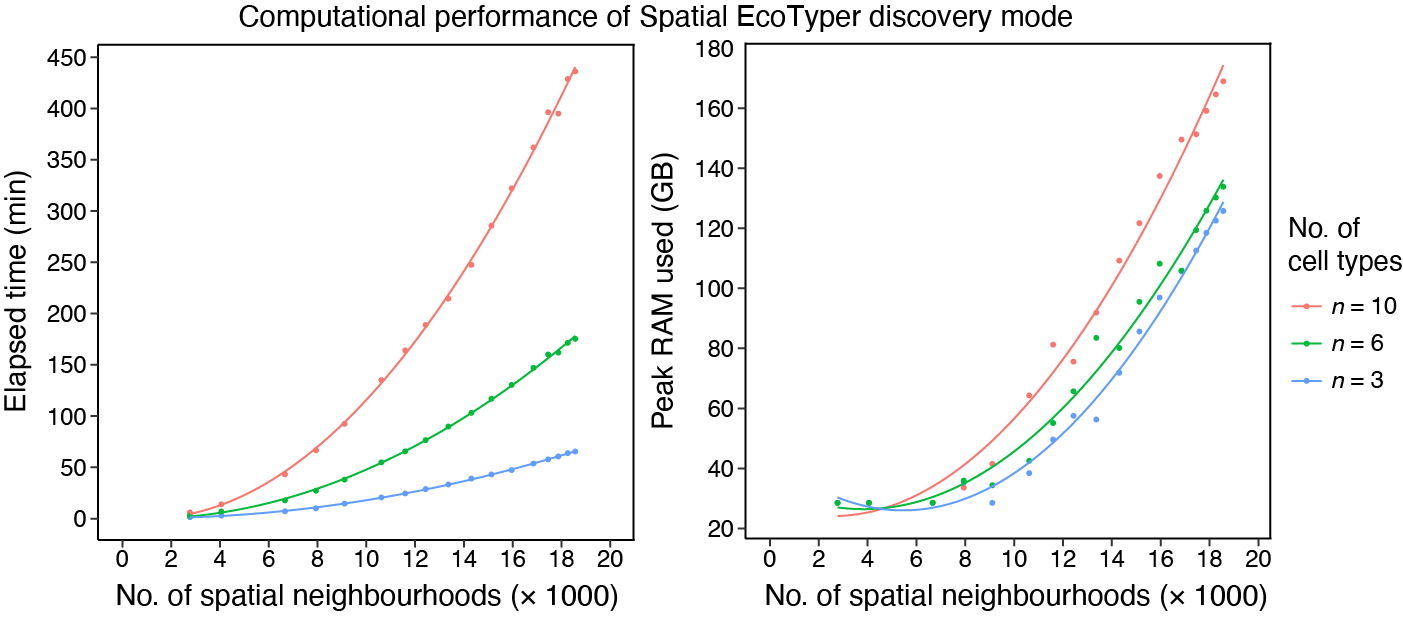 For example, an analysis involving 10 cell types and 18,564 SNs required
436 minutes of runtime and reached a peak memory usage of 169 GB. This
benchmark was obtained on a computing cluster equipped with an AMD EPYC
7543 processor (2.75 GHz) and 256 GB RAM, using a single CPU core and
excluding data loading time.
For example, an analysis involving 10 cell types and 18,564 SNs required
436 minutes of runtime and reached a peak memory usage of 169 GB. This
benchmark was obtained on a computing cluster equipped with an AMD EPYC
7543 processor (2.75 GHz) and 256 GB RAM, using a single CPU core and
excluding data loading time.
To reduce runtime, users can increase the number of cores via the
ncores parameter. However, this parallelization comes at
the cost of increased memory consumption.
When memory resources are limited, users can reduce the number of
spatial neighborhoods by increasing grid.size, which makes
the spatial grid coarser. Increasing radius can also reduce
the number of unique SNs, but it changes spatial resolution and should
be used carefully.
Output results
All results will be saved in the specified directory
(outdir). The output files include:
-
(SampleName)_SpatialEcoTyper_results.rdsSpatialEcoTyperresult for each sample (ifMultiSpatialEcoTyperfunction was used). - MultiSE_integrated_final.rds A matrix representing the fused similarity matrix of spatial clusters across all samples.
- MultiSE_integrated_final_hmap.pdf A heatmap visualizing the fused similarity matrix of spatial clusters, grouped by samples and SEs.
- MultiSE_metadata_final.rds Single-cell metadata with an ‘SE’ column annotating the discovered SEs, saved as an RDS file.
-
MultiSE_metadata_final.tsv The same as
MultiSE_metadata_final.rds, but saved in TSV format. - SpatialView_SEs_by_Sample.pdf A figure showing the spatial landscape of SEs across samples.
- BarView_SEs_Sample_Frac.pdf A bar plot showing the fraction of cells from each sample represented within each SE.
- BarView_SEs_Region_Frac_Avg.pdf A bar plot showing the average fraction of regions represented within each SE.
- BarView_SEs_CellType_Frac_Avg.pdf A bar plot showing the average cell type composition within each SE.
-
MultiSE_NMF_results.rds The NMF result derived from
the
nmfClusteringfunction. If a single rank was provided, it contains anNMFfitX1object . If multiple ranks were provided, it is alistcontaining the optimal number of communities (bestK), a list ofNMFfitX1objects (NMFfits), and aggplotobject (p) displaying the cophenetic coefficient for different cluster numbers. - MultiSE_NMF_Cophenetic_Dynamics.pdf A figure showing the cophenetic coefficients for different numbers of clusters, only available when multiple ranks were provided.
Identifying a different number of spatial ecotypes
SpatialEcoTyper uses NMF to group spatial clusters from multiple samples into conserved SEs. If you want to try a different number of clusters for spatial ecotype identification, you can use the nmfClustering function.
This analysis requires the fused similarity network matrix
(MultiSE_integrated_final.rds) generated from previous
step.
## Fused similarity matrix of spatial clusters across all samples
outdir = "SpatialEcoTyper_results/"
integrated = readRDS(file.path(outdir, "MultiSE_integrated_final.rds"))
dim(integrated)## [1] 385 385
## Single-cell metadata with an 'SE' column annotating the discovered SEs
# Column InitSE: Sample-level spatial clusters (before integration)
# Column SE: Conserved spatial ecotypes (after cross-sample integration)
finalmeta = readRDS(file.path(outdir, "MultiSE_metadata_final.rds"))
head(finalmeta[, c("Sample", "X", "Y", "CellType", "Region", "InitSE", "SE")])## Sample X Y CellType Region
## HumanMelanomaPatient1__cell_3655 SKCM 1894.706 -6367.766 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3657 SKCM 1942.480 -6369.602 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3658 SKCM 1963.007 -6374.026 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3660 SKCM 1981.600 -6372.266 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3661 SKCM 1742.939 -6374.851 Fibroblast Stroma
## HumanMelanomaPatient1__cell_3663 SKCM 1921.683 -6383.309 Fibroblast Stroma
## InitSE SE
## HumanMelanomaPatient1__cell_3655 SKCM..InitSE69 SE7
## HumanMelanomaPatient1__cell_3657 SKCM..InitSE146 SE1
## HumanMelanomaPatient1__cell_3658 SKCM..InitSE146 SE1
## HumanMelanomaPatient1__cell_3660 SKCM..InitSE146 SE1
## HumanMelanomaPatient1__cell_3661 SKCM..InitSE92 SE7
## HumanMelanomaPatient1__cell_3663 SKCM..InitSE69 SE7For instance, if you want to identify 10 SEs, you can specific
ranks = 10 to identify 10 clusters.
nmf_res <- nmfClustering(integrated, ranks = 10, nrun.per.rank = 30,
seed = 1, ncores = 8)
ses <- predict(nmf_res)
## Add the newly defined SEs into the meta data
finalmeta$SE = paste0("SE", ses[finalmeta$InitSE])
head(finalmeta)Determining the optimal number of clusters
To determine the optimal number of clusters, you can provide multiple
candidate values to the ranks argument in the nmfClustering function, which
evaluates each specified rank and computes the corresponding cophenetic
coefficient to assess clustering stability. Running time for this step
could be time-consuming, and the time requirement is dependent on the
data size, the number of ranks to test, the number of runs per rank, and
the number of cores for parallel analysis.
nmf_res <- nmfClustering(integrated, ranks = 2:30,
nrun.per.rank = 30, # Recommend 30 or higher
min.coph = 0.95,
ncores = 4,
seed = 1)
paste0("The selected rank is ", nmf_res$bestK)
## Visualize the change of cophenetic coefficient across ranks
plot(nmf_res$p)Once the optimal rank is determined, you can use it to generate the final SE annotations.
Visualizing SEs
Visualizing the integrated similarity matrix
After updating the NMF clustering results, you can re-order spatial clusters across samples and visualize the fused similarity matrix on a heatmap.
# Order SN clusters by SEs
tmpmeta = finalmeta %>% filter(!is.na(SE)) %>% arrange(SE) %>%
distinct(InitSE, .keep_all = TRUE) %>% as.data.frame
rownames(tmpmeta) = tmpmeta$InitSE
integrated = integrated[tmpmeta$InitSE, tmpmeta$InitSE]
# Annotation
ann <- tmpmeta[, c("Sample", "SE")]
SE_cols <- getColors(length(unique(ann$SE)), palette = 1) # colors for SEs
names(SE_cols) <- unique(ann$SE)
sample_cols <- getColors(length(unique(ann$Sample)), palette = 2) # colors for samples
names(sample_cols) <- unique(ann$Sample)
## draw heatmap
p = HeatmapView(integrated, show_row_names = FALSE, show_column_names = FALSE,
top_ann = ann, top_ann_col = list(Sample = sample_cols, SE = SE_cols))
p = ComplexHeatmap::draw(p)
drawRectangleAnnotation(p, ann$SE, ann$SE)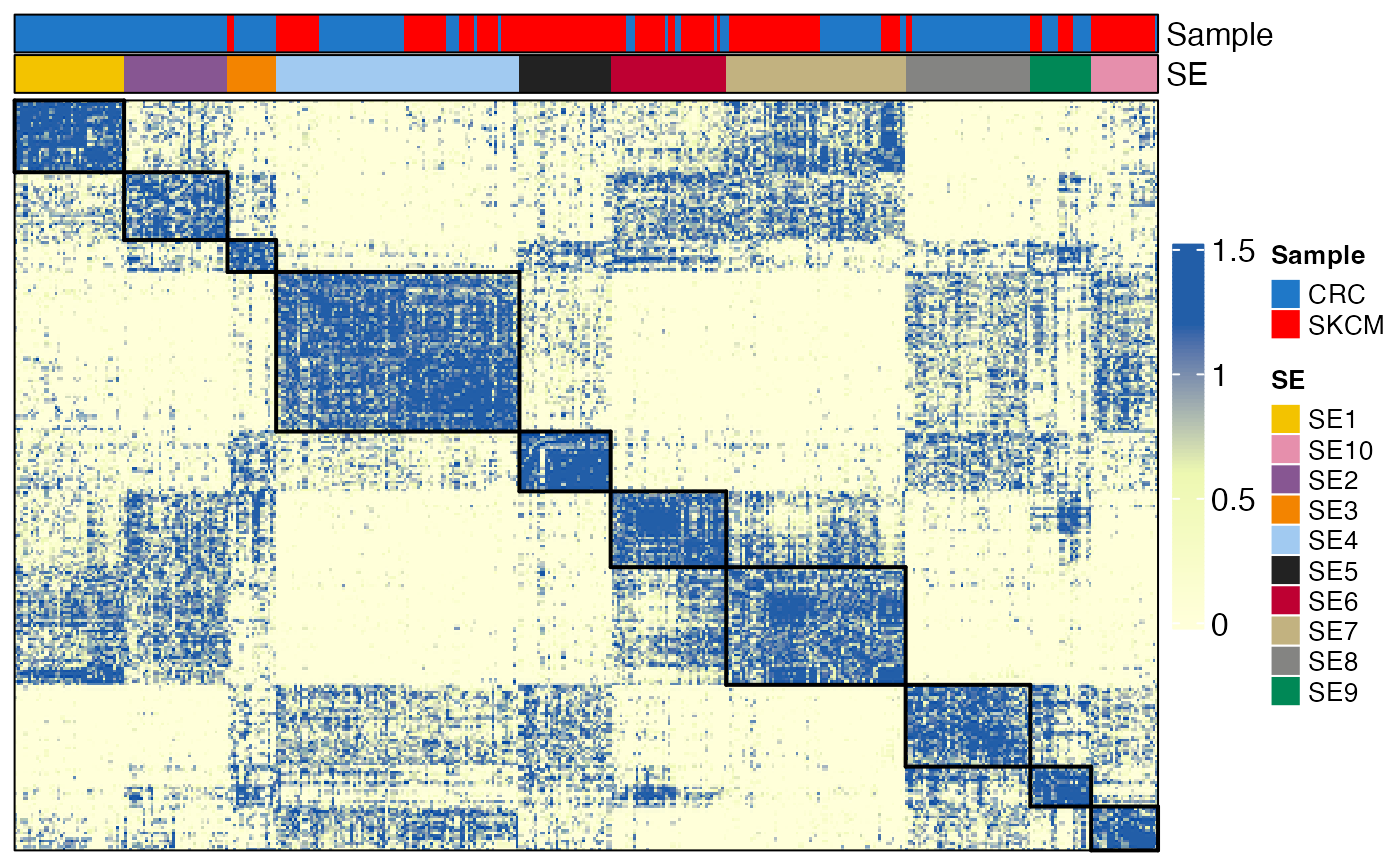
Visualizing SEs in the tissue
The spatial distribution of SEs within the tissue can be visualized using the SpatialView function.
SpatialView(finalmeta, by = "SE") + facet_wrap(~Sample, scales = "free")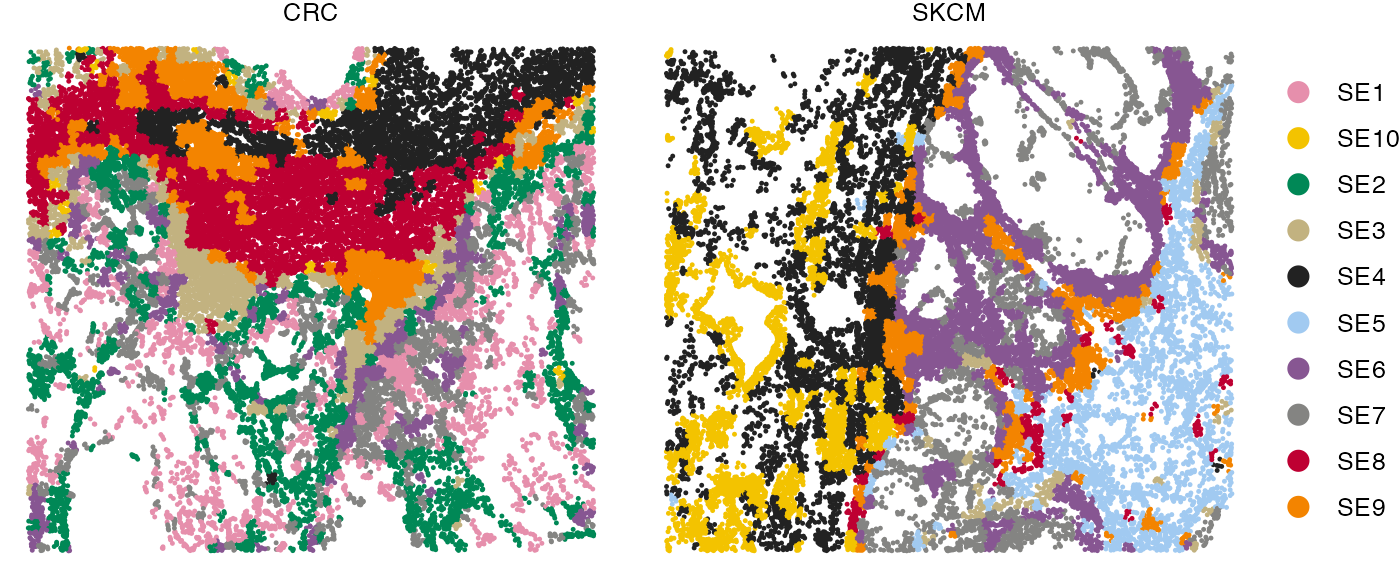
Association between SEs and pre-annotated regions
This bar plot shows the enrichment of SEs in pre-defined regions (e.g., tumor and stroma).
gg <- finalmeta %>% filter(!is.na(SE)) %>% count(SE, Region, Sample) %>%
group_by(Sample, SE) %>% mutate(Frac = n / sum(n)) %>% ## cell type fractions within each sample
group_by(SE, Region) %>% summarise(Frac = mean(Frac)) ## average cell type fractions across all samples
ggplot(gg, aes(SE, Frac, fill = Region)) +
geom_bar(stat = "identity", position = "fill") +
scale_fill_brewer(type = "qual", palette = "Set1") +
theme_bw(base_size = 14) + coord_flip() +
labs(y = "Fraction")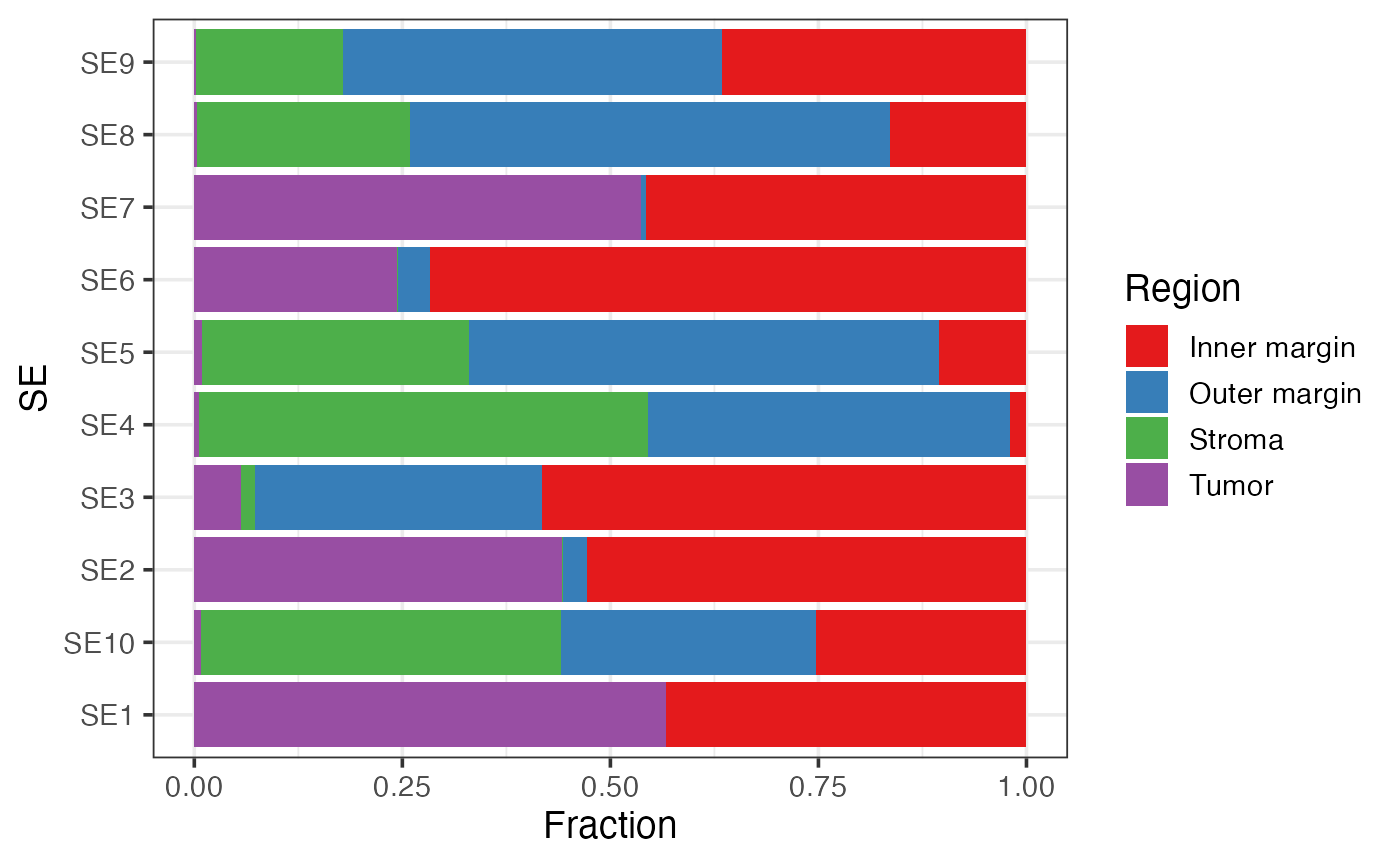
Association between SEs and pre-annotated regions within each sample
gg <- finalmeta %>% filter(!is.na(SE)) %>% count(SE, Region, Sample)
ggplot(gg, aes(SE, n, fill = Region)) +
geom_bar(stat = "identity", position = "fill") +
scale_fill_brewer(type = "qual", palette = "Set1") +
facet_wrap(~Sample) +
theme_bw(base_size = 14) + coord_flip() +
labs(y = "Fraction of regions")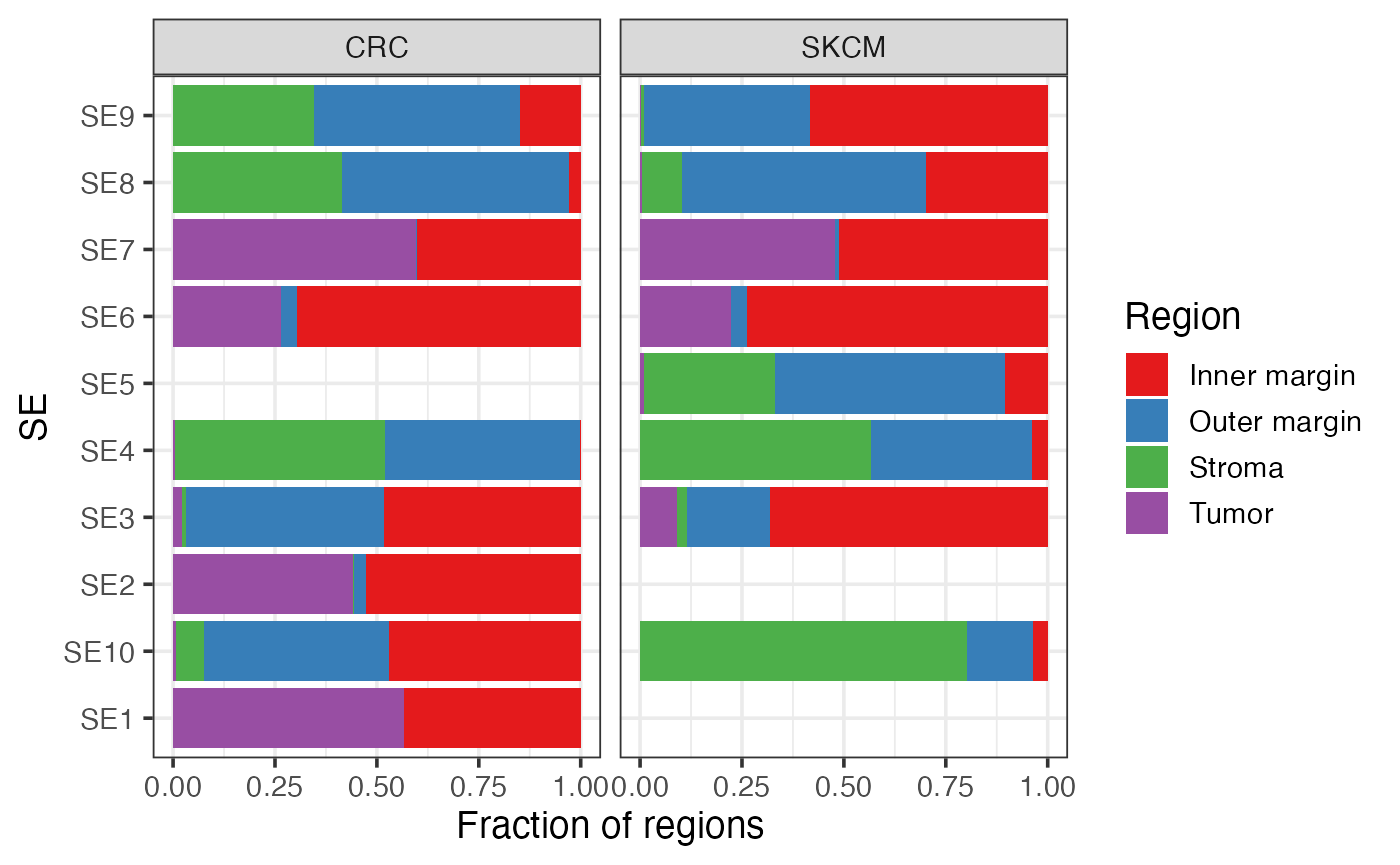
Cell type composition of SEs
We can visualize the cell type composition of SEs in a bar plot. The cell type fractions can be averaged across the two samples.
Average cell type composition of SEs across all samples
gg <- finalmeta %>% filter(!is.na(SE)) %>% count(SE, CellType, Sample) %>%
group_by(Sample, SE) %>% mutate(Frac = n / sum(n)) %>% ## cell type fractions within each sample
group_by(SE, CellType) %>% summarise(Frac = mean(Frac)) ## average cell type fractions across all samples
ggplot(gg, aes(SE, Frac, fill = CellType)) +
geom_bar(stat = "identity", position = "fill") +
scale_fill_manual(values = pals::cols25()) +
theme_bw(base_size = 14) + coord_flip() +
labs(y = "Cell type abundance")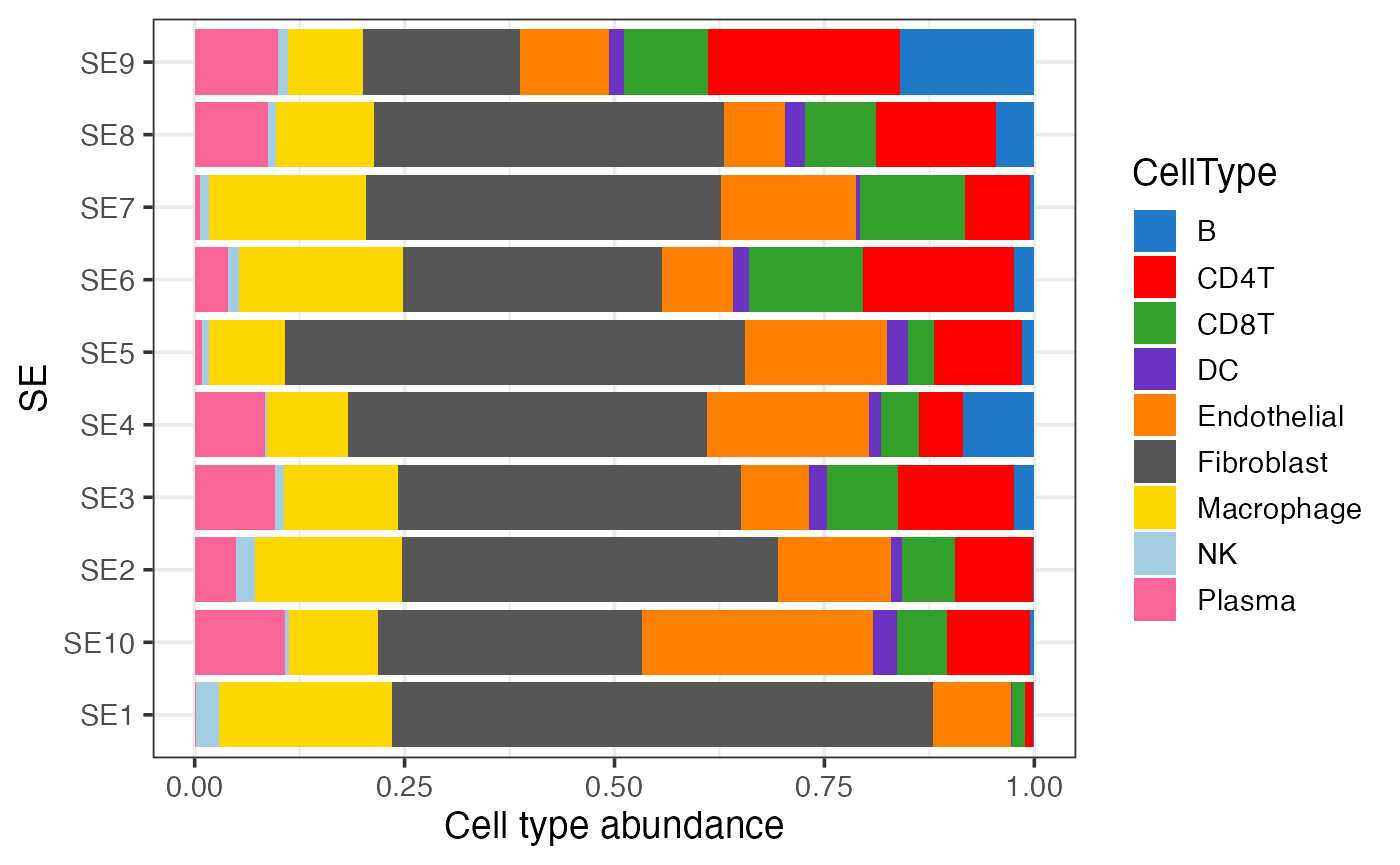
Cell type composition of SEs within each sample
gg <- finalmeta %>% filter(!is.na(SE)) %>% count(SE, CellType, Sample)
ggplot(gg, aes(SE, n, fill = CellType)) +
geom_bar(stat = "identity", position = "fill") +
scale_fill_manual(values = pals::cols25()) +
facet_wrap(~Sample) +
theme_bw(base_size = 14) + coord_flip() +
labs(y = "Cell type abundance")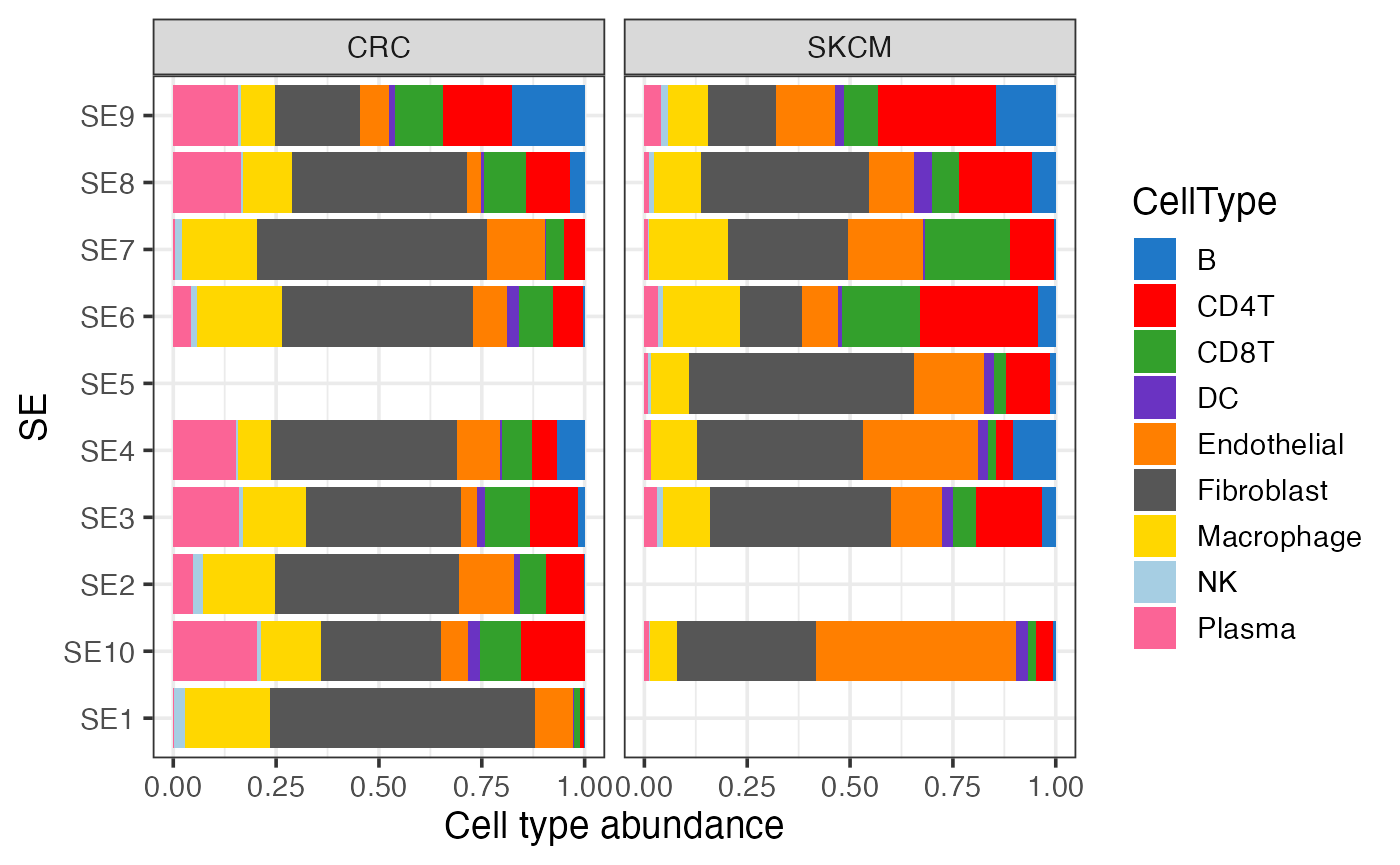
Distance of SEs to tumor/stroma interface
This box plot visualizes the distribution of distances of SEs to the tumor/stroma interface. Positive distances indicate cells located within the tumor region, while negative distances denote cells within the stroma. The SEs are ordered by their median distance, highlighting their spatial localization relative to the tumor/stroma interface.
gg <- finalmeta %>% filter(!is.na(SE)) %>% group_by(Sample, SE) %>%
summarise(Dist2Interface = mean(Dist2Interface)) %>% arrange(Dist2Interface)
gg$SE = factor(gg$SE, levels = unique(gg$SE))
ggplot(gg, aes(SE, Dist2Interface)) + geom_boxplot() +
geom_point(aes(color = Sample)) + theme_bw() +
labs(y = "Distance to tumor/stroma interface (μm)")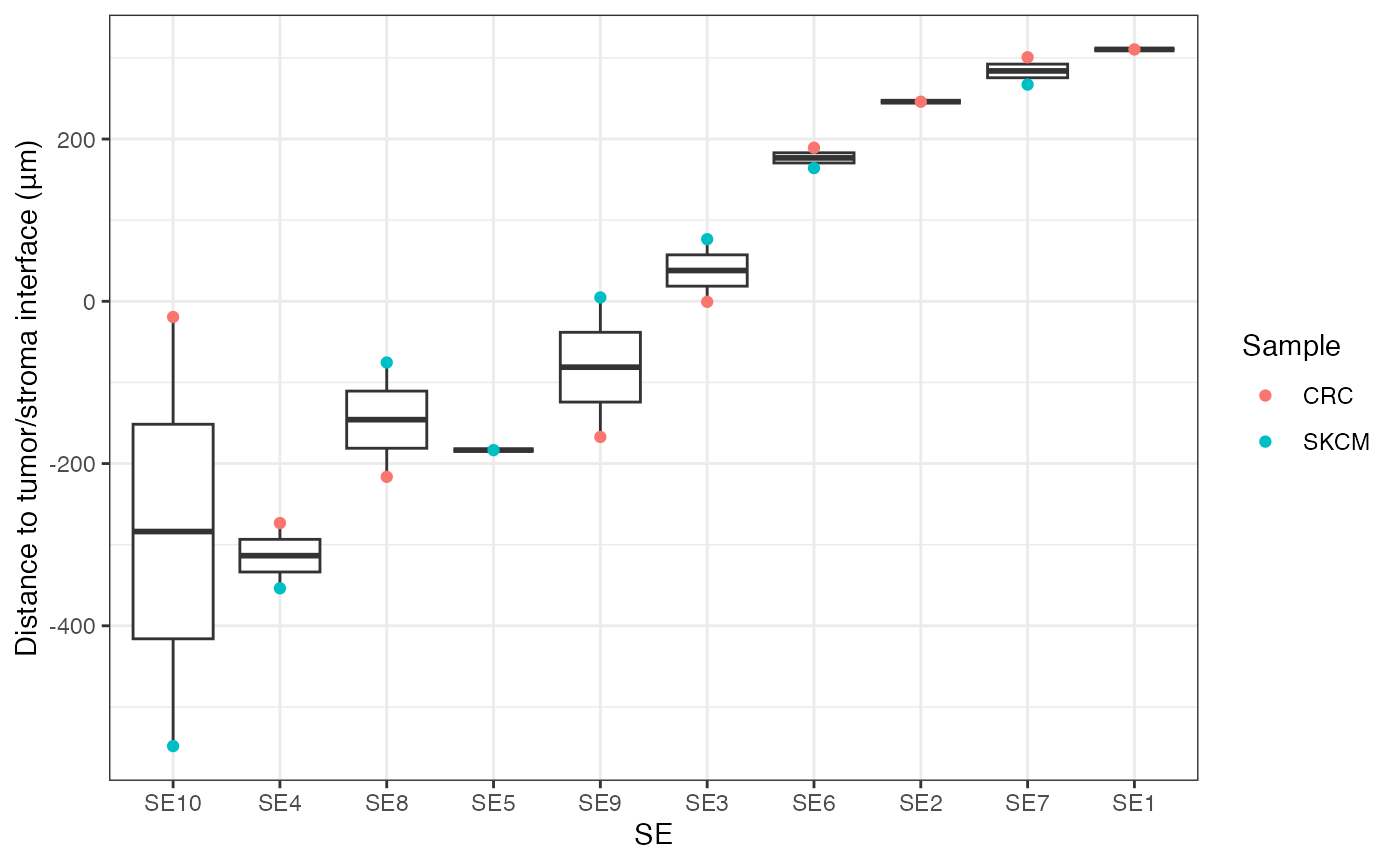
Identification of cell-type-specific SE markers
You can identify cell-type-specific SE markers by differential
expression analysis using the presto
package. Below is an example of how to identify fibroblast-specific
markers for each SE.
DE analysis within each sample
require("presto")
## DE analysis within the first sample
tmpmeta1 = finalmeta %>% filter(CellType=="Fibroblast" & Sample=="SKCM" & (!is.na(SE)))
tmpdata1 = normdata1[, tmpmeta1$CID]
degs1 = wilcoxauc(tmpdata1, tmpmeta1$SE)
## DE analysis within the second sample
tmpmeta2 = finalmeta %>% filter(CellType=="Fibroblast" & Sample=="CRC" & (!is.na(SE)))
tmpdata2 = normdata2[, tmpmeta2$CID]
degs2 = wilcoxauc(tmpdata2, tmpmeta2$SE) # DE analysis
head(degs1)## feature group avgExpr logFC statistic auc pval
## 1 PDK4 SE1 0.65754094 0.24518554 1991907 0.5279472 1.817392e-04
## 2 TNFRSF17 SE1 0.00000000 0.00000000 1886393 0.4999811 9.988846e-01
## 3 ICAM3 SE1 0.00000000 0.00000000 1886393 0.4999811 9.988846e-01
## 4 FAP SE1 0.41089988 -0.34706947 1704947 0.4518894 3.225772e-07
## 5 GZMB SE1 0.02402809 -0.05817573 1856047 0.4919380 1.610637e-02
## 6 TSC2 SE1 0.02312765 -0.02541757 1871793 0.4961114 1.452452e-01
## padj pct_in pct_out
## 1 7.766634e-04 16.2271805 11.328891
## 2 9.988846e-01 0.0000000 0.000000
## 3 9.988846e-01 0.0000000 0.000000
## 4 1.966934e-06 10.9533469 20.841500
## 5 5.229339e-02 0.6085193 2.221351
## 6 3.802231e-01 0.6085193 1.385078
head(degs2)## feature group avgExpr logFC statistic auc pval
## 1 PDK4 SE1 0.5768008 -0.16802819 4118585 0.4790122 1.392025e-02
## 2 CCL26 SE1 0.0000000 0.00000000 4298863 0.4999794 9.986864e-01
## 3 CX3CL1 SE1 0.4482779 0.20030746 4523230 0.5260745 2.108270e-06
## 4 PGLYRP1 SE1 0.0000000 0.00000000 4298863 0.4999794 9.986864e-01
## 5 CD4 SE1 0.0000000 0.00000000 4298863 0.4999794 9.986864e-01
## 6 SNAI2 SE1 1.3119381 0.04667876 4349122 0.5058248 5.730139e-01
## padj pct_in pct_out
## 1 7.175389e-02 14.98195 19.085052
## 2 9.986864e-01 0.00000 0.000000
## 3 2.849014e-05 11.91336 6.759021
## 4 9.986864e-01 0.00000 0.000000
## 5 9.986864e-01 0.00000 0.000000
## 6 9.986864e-01 32.12996 31.990979Identifying conserved markers across samples
To identify markers conserved across different samples, we conduct a meta-analysis by averaging the log2 fold changes (log2FC) from the DE analyses.
library(tidyr)
degs <- merge(degs1[, c(1,2,4)], degs2[, c(1,2,4)], by = c("feature", "group"))
degs$AvgLogFC = (degs$logFC.x + degs$logFC.y) / 2
lfcs = degs %>% pivot_wider(id_cols = feature, names_from = group,
values_from = AvgLogFC) %>% as.data.frame
rownames(lfcs) <- lfcs$feature
lfcs <- lfcs[, -1]
head(lfcs)## SE1 SE2 SE3 SE4 SE5
## ACKR3 0.23129002 -0.37521116 -0.69644369 -0.20866491 -0.03791532
## ACTA2 2.26183640 -0.09824403 0.09400553 -0.22370086 -0.18170887
## ADAMTS4 -0.04167099 0.63380839 0.78192961 0.20039317 -0.28792436
## AKT1 0.15759620 1.08038975 0.95750771 0.05349286 -0.39196038
## AKT2 -0.03270727 0.02103533 0.11173328 -0.08599313 -0.18438298
## AKT3 0.43349780 0.35193498 0.10164257 -0.19753143 -0.28513940
## SE6 SE7 SE8 SE9
## ACKR3 -0.292968737 0.788662831 0.66468166 0.09324492
## ACTA2 0.797982497 -0.051484034 -0.82402411 0.18951480
## ADAMTS4 0.066685517 -0.608592924 0.01962479 0.11986800
## AKT1 0.246800126 -0.568748749 -0.28703024 -0.23328946
## AKT2 0.008436113 0.043805780 0.06123596 0.08893108
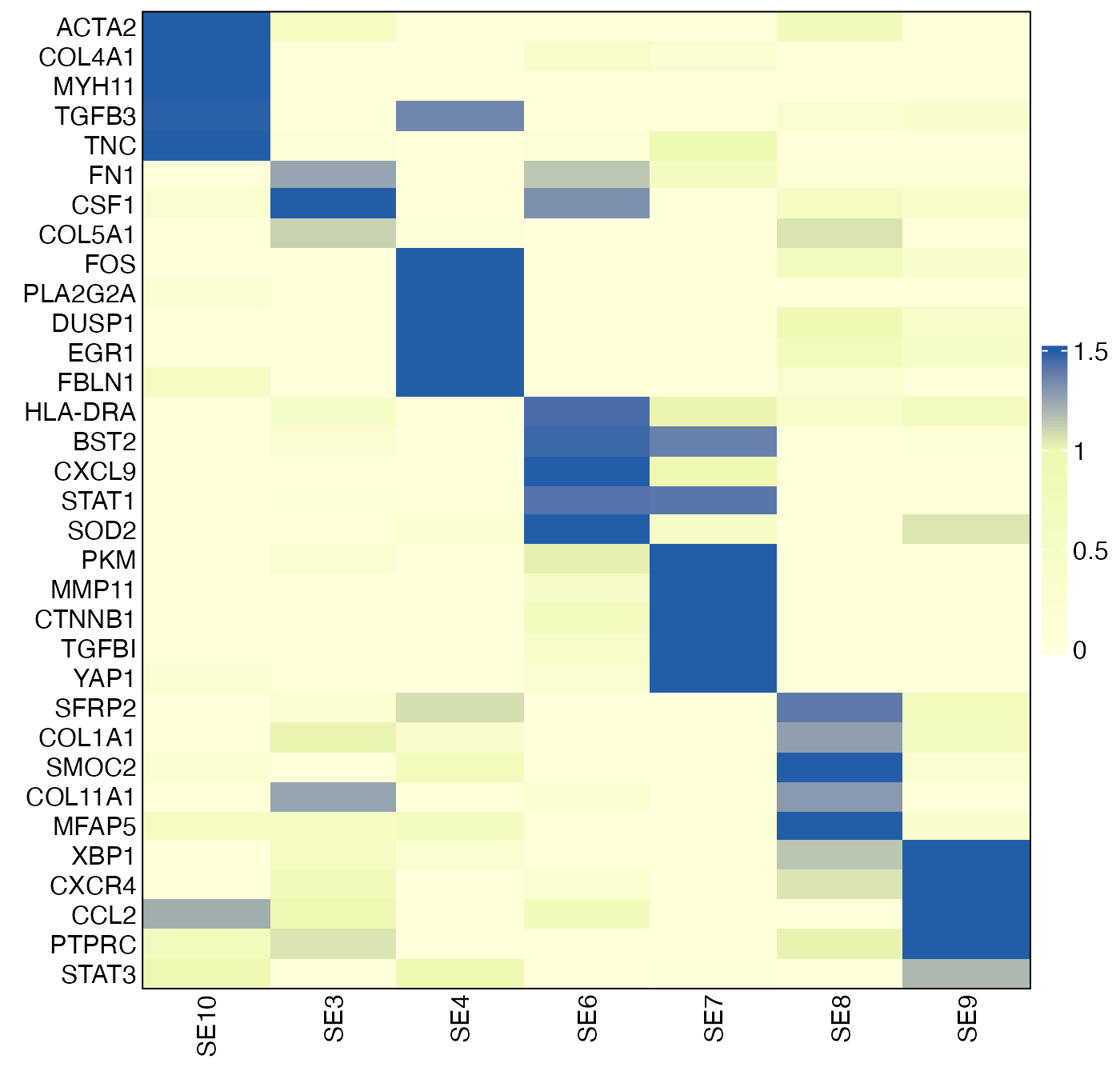
## AKT3 -0.033547946 0.007479543 0.01699382 0.28178934To identify markers that are specific to each SE-associated cell state, we calculate the difference between the maximum log2FC for each gene and the second-highest log2FC:
secondmax = apply(lfcs, 1, function(x){ -sort(-x)[2] })
delta = lfcs - secondmax
idx = delta>0 & lfcs>0.05
markers = lapply(colnames(lfcs), function(se){
gs = rownames(idx)[idx[, se]]
gs = gs[order(-lfcs[gs, se])]
gs
})
names(markers) = colnames(lfcs)
markers = markers[lengths(markers)>0]
## Select the top five markers
top5 = lapply(markers, function(x){ x[1:min(5, length(x))] })
top5## $SE1
## [1] "ACTA2" "COL4A1" "MYH11" "TNC" "ITGA1"
##
## $SE2
## [1] "CDK2" "MYC" "CTNNB1" "YAP1" "BCL2L1"
##
## $SE3
## [1] "MMP11" "BST2" "TGFBI" "STAT1" "PKM"
##
## $SE4
## [1] "CXCR4" "SOD2" "NFKBIA" "XBP1" "PTPRC"
##
## $SE5
## [1] "COL11A1" "COL1A1" "HLA-DRB1" "COL5A1" "HLA-DPA1"
##
## $SE6
## [1] "FN1" "SERPINE1" "CSF1" "CCL2" "SERPINA1"
##
## $SE7
## [1] "PLA2G2A" "FBLN1" "FOS" "DUSP1" "PDGFRA"
##
## $SE8
## [1] "TGFBR3" "PDGFRB" "TCF7L2" "CD248" "KITLG"
##
## $SE9
## [1] "GPX3" "EGFR" "FZD7" "PGF" "ANGPT2"Next step
Following spatial ecotype discovery, users can identify SE-specific cell states via leave-one-sample-out cross-validation and develop machine learning models to enable SE recovery from held-out single-cell datasets (e.g., scRNA-seq or single-cell spatial transcriptomics).
Session info
The session info allows users to replicate the exact environment and identify potential discrepancies in package versions or configurations that might be causing problems.
## R version 4.4.1 (2024-06-14)
## Platform: aarch64-apple-darwin20
## Running under: macOS 26.4.1
##
## Matrix products: default
## BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
## LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
##
## locale:
## [1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
##
## time zone: America/Los_Angeles
## tzcode source: internal
##
## attached base packages:
## [1] grid parallel stats graphics grDevices utils datasets
## [8] methods base
##
## other attached packages:
## [1] tidyr_1.3.1 presto_1.0.0 Rcpp_1.0.13
## [4] pals_1.9 SpatialEcoTyper_1.0.3 RANN_2.6.2
## [7] Matrix_1.7-0 ComplexHeatmap_2.20.0 NMF_0.28
## [10] Biobase_2.64.0 BiocGenerics_0.50.0 cluster_2.1.6
## [13] rngtools_1.5.2 registry_0.5-1 data.table_1.16.0
## [16] Seurat_5.1.0 SeuratObject_5.0.2 sp_2.1-4
## [19] ggplot2_3.5.1 dplyr_1.1.4
##
## loaded via a namespace (and not attached):
## [1] RcppAnnoy_0.0.22 splines_4.4.1 later_1.3.2
## [4] R.oo_1.26.0 tibble_3.2.1 polyclip_1.10-7
## [7] fastDummies_1.7.4 lifecycle_1.0.4 sf_1.1-0
## [10] doParallel_1.0.17 globals_0.16.3 lattice_0.22-6
## [13] MASS_7.3-60.2 magrittr_2.0.3 plotly_4.10.4
## [16] sass_0.4.9 rmarkdown_2.28 jquerylib_0.1.4
## [19] yaml_2.3.10 httpuv_1.6.15 sctransform_0.4.1
## [22] spam_2.10-0 spatstat.sparse_3.1-0 reticulate_1.39.0
## [25] mapproj_1.2.11 cowplot_1.1.3 pbapply_1.7-2
## [28] DBI_1.3.0 RColorBrewer_1.1-3 maps_3.4.2
## [31] abind_1.4-5 Rtsne_0.17 R.utils_2.12.3
## [34] purrr_1.0.2 circlize_0.4.16 IRanges_2.38.1
## [37] S4Vectors_0.42.1 ggrepel_0.9.6 irlba_2.3.5.1
## [40] listenv_0.9.1 spatstat.utils_3.1-0 units_1.0-1
## [43] goftest_1.2-3 RSpectra_0.16-2 spatstat.random_3.3-1
## [46] fitdistrplus_1.2-1 parallelly_1.38.0 pkgdown_2.1.0
## [49] leiden_0.4.3.1 codetools_0.2-20 tidyselect_1.2.1
## [52] shape_1.4.6.1 farver_2.1.2 matrixStats_1.4.1
## [55] stats4_4.4.1 spatstat.explore_3.3-2 jsonlite_1.8.8
## [58] GetoptLong_1.0.5 e1071_1.7-16 progressr_0.14.0
## [61] ggridges_0.5.6 survival_3.6-4 iterators_1.0.14
## [64] systemfonts_1.1.0 foreach_1.5.2 tools_4.4.1
## [67] ragg_1.3.2 ica_1.0-3 glue_1.7.0
## [70] gridExtra_2.3 xfun_0.52 withr_3.0.1
## [73] BiocManager_1.30.25 fastmap_1.2.0 boot_1.3-30
## [76] fansi_1.0.6 spData_2.3.4 digest_0.6.37
## [79] R6_2.5.1 mime_0.12 wk_0.9.5
## [82] textshaping_0.4.0 colorspace_2.1-1 scattermore_1.2
## [85] tensor_1.5 dichromat_2.0-0.1 spatstat.data_3.1-2
## [88] R.methodsS3_1.8.2 utf8_1.2.4 generics_0.1.3
## [91] class_7.3-22 httr_1.4.7 htmlwidgets_1.6.4
## [94] spdep_1.4-2 uwot_0.2.2 pkgconfig_2.0.3
## [97] gtable_0.3.5 lmtest_0.9-40 htmltools_0.5.8.1
## [100] dotCall64_1.1-1 clue_0.3-65 scales_1.3.0
## [103] png_0.1-8 spatstat.univar_3.0-1 knitr_1.48
## [106] rstudioapi_0.16.0 reshape2_1.4.4 rjson_0.2.22
## [109] nlme_3.1-164 proxy_0.4-27 cachem_1.1.0
## [112] zoo_1.8-12 GlobalOptions_0.1.2 stringr_1.5.1
## [115] KernSmooth_2.23-24 miniUI_0.1.1.1 s2_1.1.9
## [118] desc_1.4.3 pillar_1.9.0 vctrs_0.6.5
## [121] promises_1.3.0 xtable_1.8-4 evaluate_0.24.0
## [124] magick_2.8.5 cli_3.6.3 compiler_4.4.1
## [127] rlang_1.1.4 crayon_1.5.3 future.apply_1.11.2
## [130] labeling_0.4.3 classInt_0.4-11 plyr_1.8.9
## [133] fs_1.6.4 stringi_1.8.4 viridisLite_0.4.2
## [136] deldir_2.0-4 gridBase_0.4-7 munsell_0.5.1
## [139] lazyeval_0.2.2 spatstat.geom_3.3-2 RcppHNSW_0.6.0
## [142] patchwork_1.2.0 future_1.34.0 shiny_1.9.1
## [145] highr_0.11 ROCR_1.0-11 igraph_2.0.3
## [148] bslib_0.8.0